Principles
Metabolic Imaging by NAD(P)H and FAD FLIM
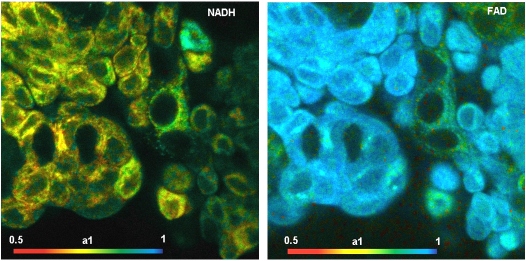
NAD(P)H (nicotinamide adenine (pyridine) dinucleotide) and FAD (flavin adenine dinucleotide) are coencymes involved in the cell metabolism. Both NAD(P)H and FAD are fluorescent. FAD and, especially, NAD(P)H are unique in the sense that their fluorescence intensities and fluorescence decay functions bear direct information on the metabolic state of the cells: The fluorescence lifetimes of NAD(P)H and FAD depend on the binding to proteins. Unbound NAD(P)H has a fluorescence lifetime of about 0.3 to 0.4 ns. Bound NAD(P)H has a lifetime of about 1.2 ns [9]. For FAD the effect of binding is opposite: Bound FAD has a lifetime of a few 100 ps, unbound FAD of a few ns.
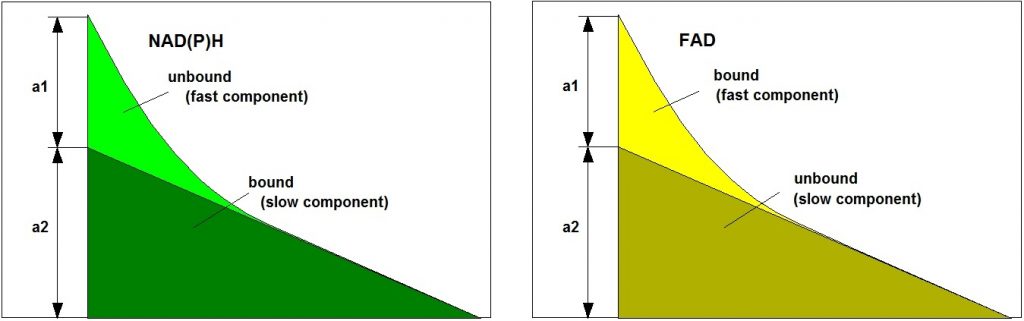
Importantly, the ratios of the amounts of bound and unbound NAD(P)H and of bound and unbound FAD depend on the type of the metabolism. A shift from glycolysis to oxidative phosphorylation or back results in a change in the unbound/bound ratios: Glycolysis yields high a1/a2 for NAD(P)H and low a1/a2 for FAD, oxydative phosphorylation yields low a1/a2 for NAD(P)H and high a1/a2 for FAD. The bound/unbound ratios directly reflect the ‘Warburg Effect’: In normal cells oxidative phosphorylation dominates, in cancer cells reductive glycolysis. Normal cells and cancer cells can therefore be distinguished by their a1/a2 ratios, or by a1 alone (because a1+a2=1). It should be noted that changes in a1 or a1/a2 are implicitly present also in the amplitude-weighted lifetime, tm and the intensity-weighted lifetime, ti. However, the lifetimes of the decay components themselves are influenced by other molecular-environment parameters, such as mitochondrial pH. The lifetime can therefore not be used as an absolute discrimination parameter.
Excitation and Emission Wavelengths
Approximate excitation and emission spectra of NAD(P)H and FAD are shown in the figure below. The figure shows that the fluorescence signals from NAD(P)H and FAD can only be separated if different excitation wavelengths and different detection wavelengths are used. The technique of choice is FLIM with laser-wavelength multiplexing, as it is implemented in the bh DCS-120 FLIM systems.
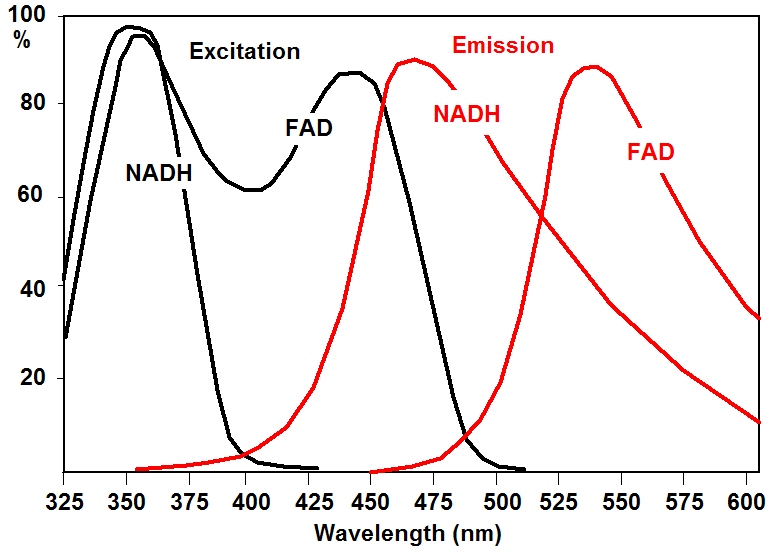
Suitable one-photon excitation wavelengths are 350 to 375 nm for NADH and 405 to 420 nm for FAD. Two photon-excitation wavelengths are 750 nm and 920 nm, respectively. Detection wavelength intervals are about 425 to 475 nm for NAD(P)H and 475 to 600 nm for FAD.
Simultaneous Measurement of NAD(P)H and FAD
To minimise the influence of photobleaching, focus drift, an possible physiological changes it is desirable to record the FLIM data of NAD(P)H and FAD quasi simultaneously. This can be obtained by laser multiplexing and multiplexed TCSPC: The lasers are multiplexed synchronously with the scanner, either line by line or frame by frame. The signals in the two emission wavelength intervals are recorded by two parallel FLIM channels. All the required functions are implemented in the bh DCS-120 confocal scanning FLIM system. Metabolic FLIM with the DCS-120 is thus only a matter of using the right lasers and choosing the right setup parameters. With bh FLIM systems attached to other laser scanning microscopes metabolic FLIM can, in principles be performed as well if the right lasers are used and are prepared for multiplexing.
Typical Results
The images shown below were recorded with a DCS-120 system with BDL-SMN lasers of 375 nm and 405 nm wavelength. The lasers were multiplexed frame by frame, and the images recorded in the two channels of the DCS-120 system.
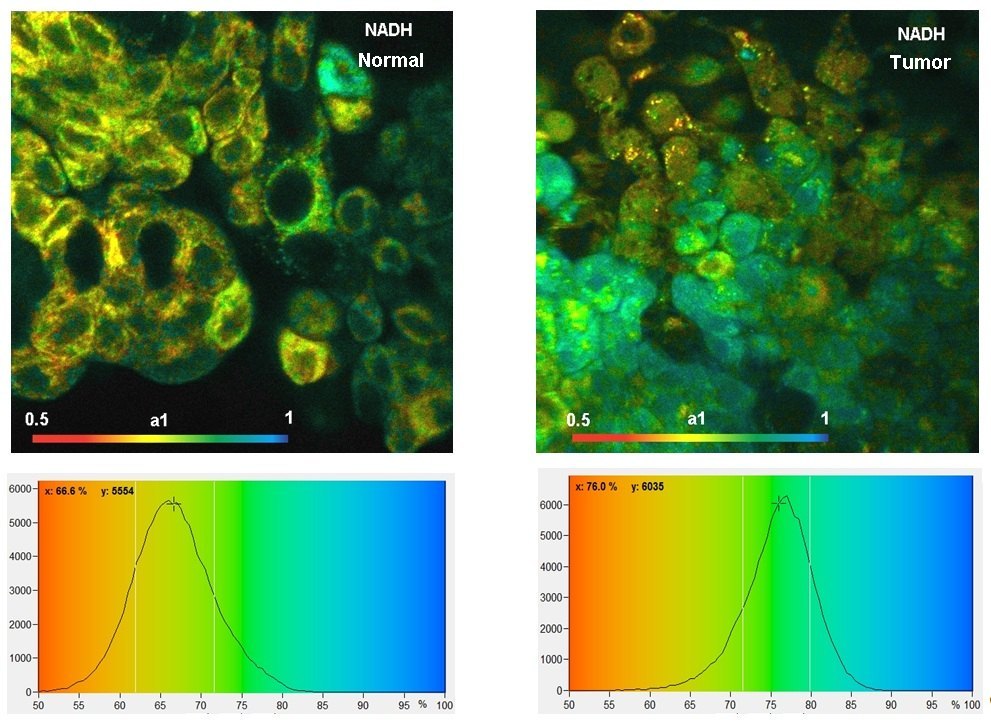
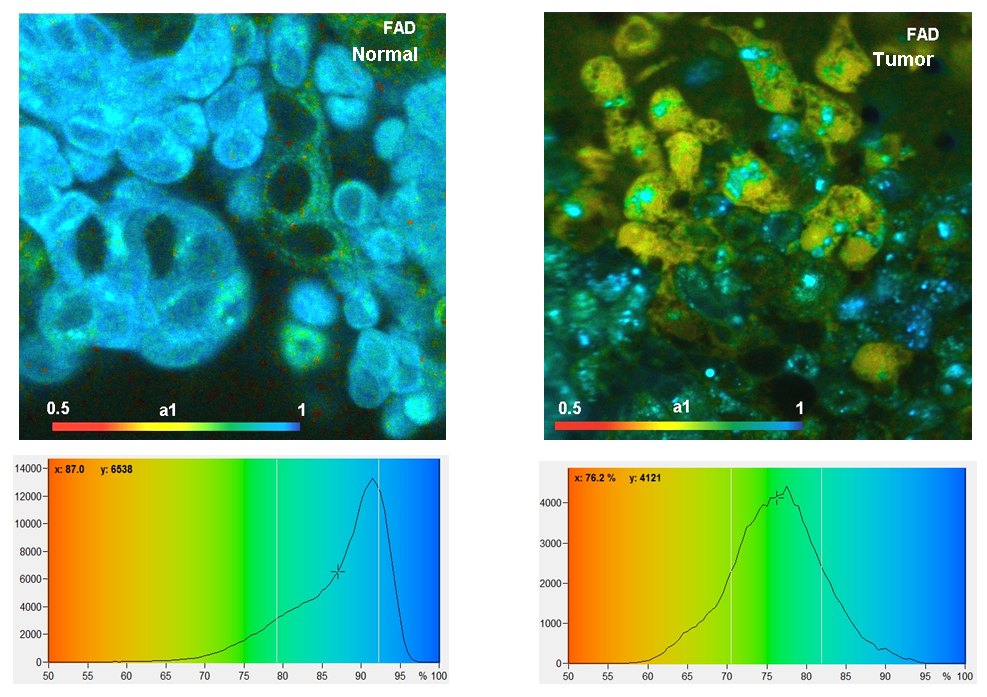
The data shown above demonstrate show that the distribution of the amplitude of the fast decay component, a1, in the NAD(P)H and in the FAD images are different for normal cells and tumor cells. For more information please see bh TCSPC Handbook and application note ‘Metabolic Imaging with the DCS-120 Confocal FLIM System: Simultaneous FLIM of NAD(P)H and FAD’.
Contribution of FMN to FAD Data
FMN is a FAD-like molecule with virtually identical excitation and emission wavelength but longer fluorescence lifetime. However, FMN does not show the same metabolism-dependent change in its fluorescence lifetime. If metabolic parameters are to be derived from FAD quantitatively the contribution from FNM must be taken into account. With the high time resolution and recording acuracy of the bh FLIM systems the decay times and amplitudes of bound FAD, free FAD, and FMN can be split by SPCImage NG using triple-exponential decay analysis. Please see application note ‘DCS-120 FLIM System Detects FMN in Live Cells’. An example is shown in the figure below.

References
References Related to Metabolic Imaging
For more references please see W. Becker, The bh TCSPC Handbook.
- R. Alam, H. Wallrabe, Z. Svindrych, A. K. Chaudhary, K. G. Christopher, D. Chandra, A. Periasamy, Investigation of Mitochondrial Metabolic Response to Doxorubicin in Prostate Cancer Cells: An NADH, FAD and Tryptophan FLIM Assay. Scientific Reports 7 (2017)
- Awasthi, T. Nakabayashi, L. Li, N. Ohta, Effects of Nanosecond Pulsed Electric Field on Intracellular NADH Autofluorescence: A Comparison between Normal and Cancer Cells. ACS Omega 2, 2916-2924 (2017)
- W. Becker, R. Suarez-Ibarrola, A. Miernik, L. Braun, Metabolic Imaging by Simultaneous FLIM of NAD(P)H and FAD. Current Directions in Biomedical Engineering 5(1), 1-3 (2019
- K. Bird , L. Yan , K. M. Vrotsos , K. E. Eliceiri , E. M. Vaughan. Metabolic mapping of MCF10A human breast cells via multiphoton fluorescence lifetime imaging of coenzyme NADH. Cancer Res 65:8766–8773 (2005)
- Wolfgang Becker, Hauke Studier, Cornelia Wetzker, Ultra-fast HPM detectors improve NAD(P)H FLIM. Proc. SPIE 10498, 1049806 (2018)
- S. Blacker., M. R. Duchen, Investigating mitochondrial redox state using NADH and NADPH autofluorescence. Free Radical Biology and Medicine 100, 53–65 (2016)
- S. Blacker, Z. F. Mann, J. E. Gale, M. Ziegler, A. J. Bain, G. Szabadkai, M. R. Duchen, Separating NADH and NADPH fluorescence in live cells and tissues using FLIM. Nature Communications 5, 3936-1 to -6 (2014)
- S. Blacker, M. R.Duchen, Investigating mitochondrial redox state using NADH and NADPH autofluorescence. Free Radical Biology and Medicine 100, 53–65 (2016)
- 218. Thomas S. Blacker, Michael D. E. Sewell, Gyorgy Szabadkai, Michael R. Duchen, Metabolic Profiling of Live Cancer Tissues Using NAD(P)H Fluorescence Lifetime Imaging. In: Majda Haznadar (ed.), Cancer Metabolism: Methods and Protocols, Methods in Molecular Biology, 1928, 365-386 (2019)
- Chance, Pyridine nucleotide as an indicator of the oxygen requirements for energy-linked functions of mitochondria. Circ Res 38, I31–I38 (1976)
- Chorvat, A. Chorvatova, Spectrally resolved time-correlated single photon counting: a novel approach for characterization of endogenous fluorescence in isolated cardiac myocytes. Eur. Biophys. J. 36:73-83 (2006)
- Chorvat Jr., S. Abdulla, F. Elzwei, A. Mateasik, A. Chrovatova, Screening of cardiomyocyte fluorescence during cell contraction by multi-dimensional TCSPC. Proc. SPIE 6860, 686029-1 to -12 (2008)
- Chorvat, A. Chorvatova, Multi-wavelength fluorescence lifetime spectroscopy: a new approach to the study of endogenous fluorescence in living cells and tissues. Laser Phys. Lett. 6 175-193 (2009)
- Chorvat Jr., A. Mateasik, Y.g Cheng, N.y Poirier, J. Miro, N.S. Dahdah3, A. Chorvatova, Rejection of transplanted hearts in patients evaluated by the component analysis of multi-wavelength NAD(P)H fluorescence lifetime spectroscopy. J. Biophotonics 3 646-652 (2010)
- Chorvatova, F. Elzwiei, A. Mateasik, D. Chorvat, Effect of ouabain on metabolic oxidative state in living cardiomyocytes eavaluated by time-resolved spectroscopy of endogenous NAAD(P)H fluorescence. J. Biomed. Opt. 17(10) 101505-1 to -7 (2012)
- Chorvatova, S. Aneba, A. Mateasik, D. Chorvat Jr., B. Comte, Time-resolved fluorescence spectroscopy investigation of the effect of 4-hydroxynonenal on endogenous NAD(P)H in living cardiac myocytes. J. Biomed. Opt. 18(6), 067009-1 to -11 (2013)
- Chorvatova, A. Mateasik, D. Chorvat Jr, Spectral decomposition of NAD(P)H fluorescence components recorded by multi-wavelength fluorescence lifetime spectroscopy in living cardiac cells. Laser Phys. Lett. 10 125703-1 to -10 (2013)
- Chorvatova, Autofluorescence-assisted examination of cardiovascular system physiology and pathology. In: A. A. Heikal and V. Ghukasyan, eds., Natural Biomarkers for Cellular Metabolism: Biology, Techniques and Applications. CRC Press, Taylor & Francis Group, Boca Raton, London, New York (2015)
- Cheng, D. Chorvat, N. Poirier, J. Miro, N. Dahdah, A. Chorvatova, Spectrally and time-resolved study of NAD(P)H autofluorescence in cardiac myocytes from human biopsies. Proc. SPIE 6771, 677104-1 to -13 (2007)
- Ghukassian, F.-J. Kao, Monitoring cellular metabolism with fluorescence lifetime of reduced nicotinamide adenine dinucleotide. J. Phys. Chem. C, 113, 11532-11540 (2009)
- Ghukasyan, C.C. Hsu, C.R. Liu, F.J. Kao, T.H. Cheng, Fluorescence lifetime dynamics of enhanced green fluorescent protein aggregates with expanded polyglutamine. J. Biomed. Opt. 15(1) 016008-1 to -11 (2010)
- V. Ghukasyan, A.H. Heikal, Natural biomarkers for cellular metabolism. Biology, techniques, and applications. CRC Press, Taylor and Francis Group, Boca Raton, London, New York (2015)
- Heikal, Intracellular coenzymes as natural biomarkers for metabolic activities and mitochondrial anomalies. Biomark. Med. 4(2), 241–263 (2010)
- A. V. Izosimova, M. V. Shirmanova, V. I. Shcheslavskiy, D. A. Sachkova, A. M. Mozherov, G. V. Sharonov, E. V. Zagaynova, D. V. Yuzhakova, FLIM of NAD(P)H in Lymphatic Nodes Resolves T-Cell Immune Response to the Tumor. Int. J. Mol. Sci. 23, 5829-1 to -12 (2022)
- Kalinina, V. Shcheslavskiy, W. Becker, J. Breymayer, P. Schäfer, A. Rück, Correlative NAD(P)H-FLIM and oxygen sensing-PLIM for metabolic mapping. J. Biophotonics 9(8):800-811 (2016)
- R. Kantelhardt, J. Leppert, J. Krajewski, N. Petkus, E. Reusche, V. M. Tronnier, G. Hüttmann, A. Giese, Imaging of brain and brain tumor specimens by time-resolved multiphoton excitation microscopy ex vivo. Neuro-Onkology April 2007, 103-112
- Lukina, A. Orlova, M. Shirmanova, D. Shirokov, A. Pavlikov, A. Neubauer, H. Studier, W. Becker, E. Zagaynova, T. Yoshihara, S. Tobita, V. Shcheslavskiy, Interrogation of metabolic and oxygen states of tumors with fiber-based luminescence lifetime spectroscopy. Optics Letters 42(4) 731-734 (2017)
- Lukina, K. Yashin, E. E. Kiseleva, A. Alekseeva, Varvara Dudenkova, E. V. Zagaynova, E. Bederina, I. Medyanic, W. Becker, D. Mishra, M. Berezin, V. I. Shcheslavskiy, M. Shirmanova, Label-Free Macroscopic Fluorescence Lifetime Imaging of Brain Tumors.
- Marcek Chorvatova, Time-Resolved Spectroscopy of NAD(P)H in Live Cardiac Myocytes. In: W. Becker (ed.) Advanced time-correlated single photon counting applications. Springer, Berlin, Heidelberg, New York (2015)
- V. Meleshina, V. V. Dudenkova, M. V. Shirmanova, V. I. Shcheslavskiy, W. Becker, A. S. Bystrova, E.a I. Cherkasova, E. V. Zagaynova, Probing metabolic states of differentiating stem cells using two-photon FLIM. Scientific Reports 6, 21853-1 to 11 (2016)
- V. Meleshina, V. V. Dudenkova, A. S. Bystrova, D. S. Kuznetsova, M. V. Shirmanova, E. V. Zagaynova, Two-photon FLIM of NAD(P)H and FAD in mesenchymal stem cells undergoing either osteogenic or chondrogenic differentiation. Stem Cell Research & Therapy 8, 15-1 to -10 (2017)
- N. Pastore, H. Studier, C.S. Bonder, M.S. Roberts, Non-invasive metabolic imaging of melanoma progression. Exp. Dermatol. 26, 607–614 (2017)
- Patalay, C. Talbot, Y. Alexandrov, I. Munro, M. A. A. Neil, K. König, P. M. W. French, A. Chu, G. W. Stamp, C. Dunsby, Quantification of cellular autofluorescence of human skin using multiphoton tomography and fluorescence lifetime imaging in two spectral detection channels. Opt. Expr. 2, 3295-3308 (2011)
- D. Peltan, A.V. Thomas, I. Mikhailenko, D. K. Strickland, B. T. Hyman, C.A.F. von Arnim, Fluorescence lifetime imaging microscopy (FLIM) detects stimulus-dependent phospholylation of the low density lipoprotein receptor-related protein (LRP) in primary neurons. Biochem. Biophys. Res. Commun. 349, 24-30 (2006)
- Rück, C. Hauser, S. Mosch, S. Kalinina, Spectrally resolved fluorescence lifetime imaging to investigate cell metabolism in malignant and nonmalignant oral mucosa cells. J. Biomed. Opt. 19(9), 096005-1 to -9 (2014)
- Y. Sanchez, T.W. Prow, W.H. Sanchez, J.E.Grice, M.S. Roberts, Analysis of the metaboloic deterioration of ex-vivo skin, from ischemic necrosis, through the imaging of intracellular NAD(P)H by multiphoton tomography and fluorescence lifetime imaging microscopy (MPT-FLIM). J. Biomed. Opt. 15, 046008 (2010)
- Y. Sanchez, C. Obispo, E. Ryan, J. E. Grice, M.S. Roberts, Changes in the redox state and endogenous fluorescence of in vivo human skin due to intrinsic and phot-induced aging, measured by multiphoton tomography with fluorescence lifetime imaging. J. Biomed. Opt. 18(6), 061217-1 to -12 (2013)
- Y. Sanchez, M. Pastore, I. Haridass, K. König, W. Becker, M. S. Roberts, Fluorescence Lifetime Imaging of the Skin. In: W. Becker (ed.) Advanced time-correlated single photon counting applications. Springer, Berlin, Heidelberg, New York (2015)
- T. Sanchez, M. Venturas, S. A. Aghvami, X. Yang, S. Fraden, D. Sakkas, D. J. Needleman, Combined noninvasive metabolic and spindle imaging as potential tools for embryo and ocyte assessment. Human Reproduction, 1–13 (2019)
- T. Sanchez, T. Wang, M.Venturas Pedro, M. Zhang, E. Esencan, D. Sakkas, D. Needleman, E. Seli, Metabolic imaging with the use of fluorescence lifetime imaging microscopy (FLIM) accurately detects mitochondrial dysfunction in mouse oocytes. Reproductive Science 110, 1387-1397 (2018)
- M. Schaefer, D. Hilpert, M. Niederschweiberer, L. Neuhauser, S. Kalinina, E. Calzia, A. Rueck, B. von Einem, C.A.F. von Arnim, Mitochondrial matrix pH as a decisive factor in neurometabolic imaging. Neurophotonics 4(4):045004 (2017)
- M. Schaefer, S. Kalinina, A. Rueck, C.A.F. von Arnim, B. von Einem, NADH Autofluorescence – A Marker on its Way to Boost Bioenergetic Research. Cytometry Part A, 1-13 (2018)
- Schweitzer, S. Schenke, M. Hammer, F. Schweitzer, S. Jentsch, E. Birckner, W. Becker, Towards Metabolic Mapping of the Human Retina. Micr. Res. Tech. 70, 403-409 (2007)
- F. Sergeeva, M. V. Shirmanova, O. A. Zlobovskaya, A. I. Gavrina, V. V. Dudenkova, M. M. Lukina, K. A. Lukyanov, E. V. Zagaynova, Relationship between intracellular pH, metabolic cofactors, and caspase-3 activation in cancer cells during apoptosis. BBA – Molecular Cell Research (2017)
- J. S. Shah, M. Venturas, T. H. Sanchez, A. S. Penzias, D. J. Needleman, D. Sakkas, Fluorescence lifetime imaging microscopy (FLIM) detects differences in metabolic signatures between euploid and aneuploid human blastocysts. Human Reproduction, 37, No.3, 400–410 (2022)
- V. Shirmanova, I. N. Druzhkova, M. M. Lukina, V. V. Dudenkova, N. I. Ignatova, L. B. Snopova, V. I. Shcheslavskiy, V. V. Belousov, E. V. Zagaynova, Chemotherapy with cisplatin: insights into intracellular pH and metabolic landscape of cancer cells in vitro and in vivo. Scientific Reports 7, 8911-1 to -11
- E. A. Shirshin, M. V. Shirmanova, A. V. Gayer, M. M. Lukina, E. E. Nikonova, B. P. Yakimov, G. S. Budylin, V. V. Dudenkova, N. I. Ignatova, D. V. Komarov, V. V. Yakovlev, W. Becker, E. V. Zagaynova, V. I. Shcheslavskiy, Marian O. Scully, Label-free sensing with fluorescence lifetime imaging: The quest for matobilic heterogeneity. PNAS 119 No. 9, e2118241119 (2022)
- I. Shcheslavskiy, M. V. Shirmanova, V. V. Dudenkova, K. A. Lukyanov, A. I. Gavrina, A. V. Shumilova, E. Zagaynova, W. Becker, Fluorescence time-resolved macroimaging. Opt. Lett. 43, No. 13, 3152-5155 (2018)
- C. Skala, K. M. Riching, D. K. Bird, A. Dendron-Fitzpatrick, J. Eickhoff, K. W. Eliceiri, P. J. Keely, N. Ramanujam, In vivo multiphoton fluorescence lifetime imaging of protein-bound and free nicotinamide adenine dinucleotide in normal and precancerous epithelia. J. Biomed. Opt. 12 02401-1 to 10 (2007)
- C. Skala, K. M. Riching, A. Gendron-Fitzpatrick, J. Eickhoff, K. W. Eliceiri, J. G. White, N. Ramanujam, In vivo multiphoton microscopy of NADH and FAD redox states, fluorescence lifetimes, and cellular morphology in precancerous epithelia, PNAS 104, 19494-19499 (2007)
- C. Skala, A. Fontanella, L. Lan, J.A. Izatt, M.W. Dewhirst, Longitudinal optical imaging of tumor metabolism and hemodynamics. J. Biomed. Opt. 15(1) 011112-1 to -8 (2010)
- C. Skala, N. Ramanujam, Multiphoton Redox Ratio Imaging for Metabolic Monitoring in vivo. Methods Mol. Biol. 594, 155-162 (2010)
- Wallrabe, Z. Svindrych, S. R. Alam, K. H. Siller, T. Wang, D. Kashatus, S. Hu, A. Periasamy, Segmented cell analyses to measure redox states of autofluorescent NAD(P)H, FAD & Trp in cancer cells by FLIM. Sci. Reports 8:79, 1-11 (2018)
- M. Venturas, X. Yang, D. Sakkas, D. Needleman, Noninvasive metabolic profiling of cumulus cells, oocytes, and embryos via fluorescence lifetime imaging microscopy: a mini-review. Human Reproduction, 1–12 (2023)
- Walsh, R.C. Cook, B. Rexer, C.L. Arteaga, M.C. Skala, Optical Imaging of metabolism in HER2 overexpressing breast cancer cells. Biomedical Optics Express 3(1), 75-85 (2012)
- J. Walsh, R. S. Cook, H. C. Manning, D. J. Hicks, A. Lafontant, C. L. Arteaga, M. C. Skala, Optical Metabolic Imaging Identifies Glycolytic Levels, Subtypes, and Early-Treatment Response in Breast Cancer. Cancer Res. 73, 6164-6174 (2013)
- J. Walsh, R. S. Cook, M. E. Sanders, L. Aurisicchio, G. Ciliberto, C. L. Arteaga, M. C. Skala, Quantitative Optical Imaging of Primary Tumor Organoid Metabolism Predicts Drug Response in Breast Cancer. Cancer Res 74, OF1-OF11 (2014)
- J. Walsh, A. T. Shah, J. T. Sharick, M. C. Skala, Fluorescence Lifetime measurements of NADH in live cells and tissue. In: W. Becker (ed.) Advanced time-correlated single photon counting applications. Springer, Berlin, Heidelberg, New York (2015)
- J. Walsh, M. C. Skala, Optical metabolic imaging quantifies heterogeneous cell populations. Biomed. Opt. Expr. 6, 559-573 (2015)
- H-W. Wang, V. Ghukassyan, C.T. Chen, Y.H. Wei, H.W. Guo, J.S. Yu, F.J. Kao, Differentiation of appoptosis from necrosis by dynamic changes of reduced nicotinamide adenine dinucleotide fluorescence lifetime in live cells. J. Biomed. Opt. 13(5), 054011-1 to 9 (2008)
- Wang, R. Zhang, K. R. Bridle, A. Jayachandran, J. A. Thomas, W. Zhang, J. Yuan, Z. P. Xu, D. H. G. Crawford, X. Liang, X. Liu, M. S. Roberts, Two-photon dual imaging platform for in vivo monitoring cellularoxidative stress in liver injury. Scientific Reports 7, 45374-1 to -11 (2017)
- Yuzhakova D, Kiseleva E, Shirmanova M, Shcheslavskiy V, Sachkova D, Snopova L, Bederina E, Lukina M, Dudenkova V, Yusubalieva G, Belovezhets T, Matvienko D, Baklaushev V. Highly Invasive Fluorescent/Bioluminescent Patient-Derived Orthotopic Model of Glioblastoma in Mice. Front Oncol. 2022 Jul 13;12:897839. doi: 10.3389/fonc.2022.897839.
- W. Becker, R. Suarez-Ibarrola, A. Miernik, L. Braun, Metabolic Imaging by Simultaneous FLIM of NAD(P)H and FAD. Current Directions in Biomedical Engineering 5(1), 1-3 (2019)
- M. M. Lukina, L. E. Shimolina, N. M. Kiselev, V. E. Zagainov, D. V. Komarov, E. V. Zagaynova, M. V. Shirmanova, Interrogation of tumor metabolism in tissue samples ex vivo using fluorescence lifetime imaging of NAD(P)H. Methods Appl. Fluoresc. 8, 014002, 1-11 (2020)
- M. Lukina, K. Yashin, E. E. Kiseleva, A. Alekseeva, Varvara Dudenkova, E. V. Zagaynova, E. Bederina, I. Medyanic, W. Becker, D. Mishra, M. Berezin, V. I. Shcheslavskiy, M. Shirmanova, Label-Free Macroscopic Fluorescence Lifetime Imaging of Brain Tumors. Frontiers in Oncology 11, 666059, 1-11 (2021)
- R. Suarez-Ibarrola, L. Braun, P. Fabian Pohlmann, W. Becker, A. Bergmann, C. Gratzke, A. Miernik, K. Wilhelm, Metabolic Imaging of Urothelial Carcinoma by Simultaneous Autofluorescence Lifetime Imaging (FLIM) of NAD(P)H and FAD. Clinical Genitourinary Cancer (2020)
Products
Application Notes
- High-Resolution Measurement of NADH and FAD Fluorescence Decay with the DCS-120 MP
- Metabolic FLIM of Macroscopic Objects
- Label-Free Multiphoton FLIM of Moving Bacteria
- Metabolic Imaging with the DCS-120 Confocal FLIM System: Simultaneous FLIM of NAD(P)H and FAD
- New SPCImage Version Combines Time-Domain Analysis with Phasor Plot
